| Article ID | Journal | Published Year | Pages | File Type |
|---|---|---|---|---|
| 2039838 | Cell Reports | 2016 | 13 Pages |
•MLL-ENL has two distinct groups of downstream targets•Target gene transcription depends on either DOT1L or P-TEFb activity•Dichotomous transcription mode is consistent with separate biological function•Target gene families respond differently to pharmacological inhibitors
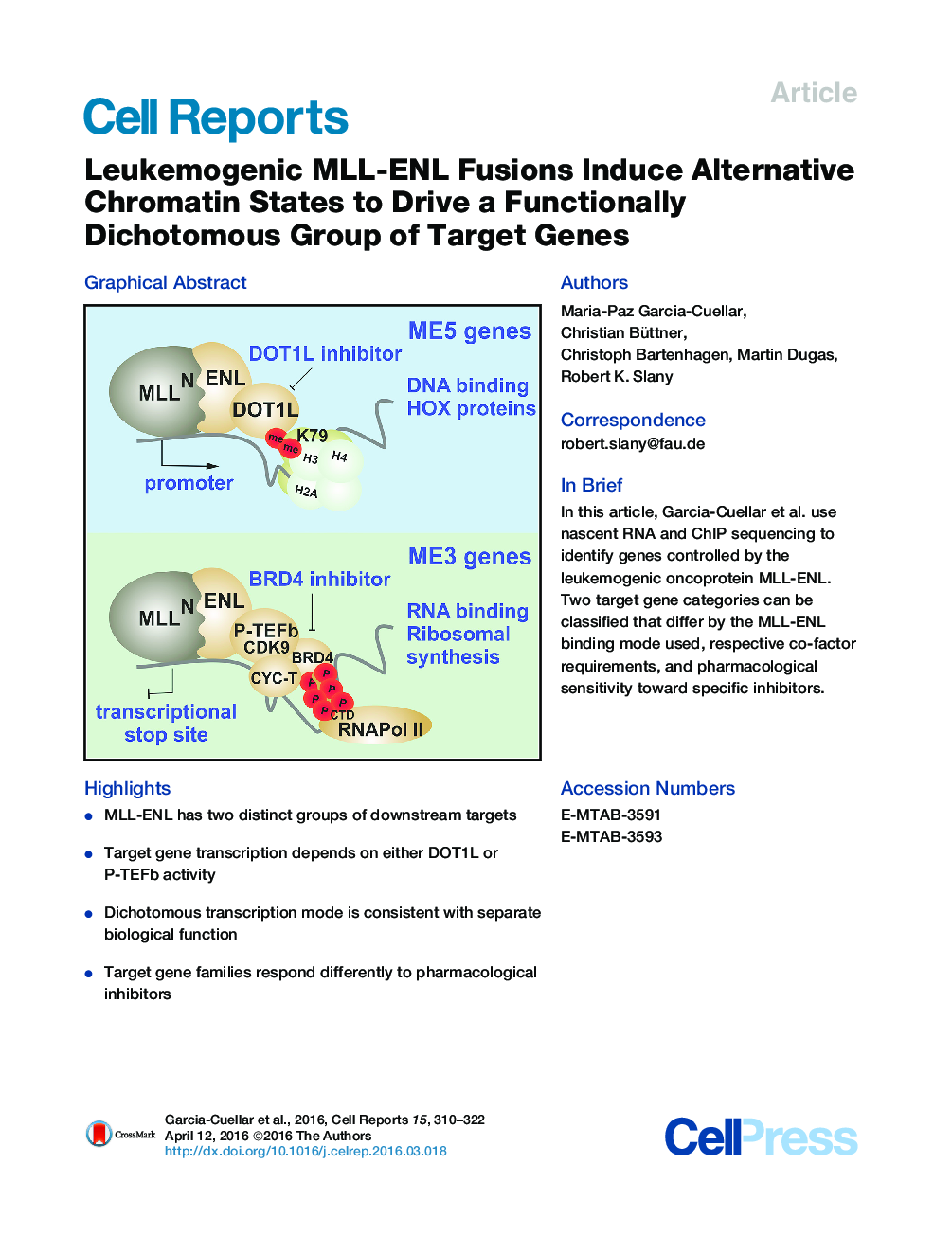
SummaryMLL fusions are leukemogenic transcription factors that enhance transcriptional elongation through modification of chromatin and RNA Pol II. Global transcription rates and chromatin changes accompanying the transformation process induced by MLL-ENL were monitored by nascent RNA-seq and ChIP-seq, revealing 165 direct target genes separated into two distinct clades. ME5 genes bound MLL-ENL at the promoter, relied on DOT1L-mediated histone methylation, and coded preferentially for transcription factors, including many homeobox genes. A distinct ME3 group accumulated MLL-ENL beyond the termination site, was dependent on P-TEFb-mediated phosphorylation of RNA Pol II for transcription, and translated mainly into proteins involved in RNA biology and ribosome assembly. This dichotomy was reflected by a differential sensitivity toward small molecule inhibitors, suggesting the possibility of a combinatorial strategy for treatment of MLL-induced leukemia.
Graphical AbstractFigure optionsDownload full-size imageDownload as PowerPoint slide