| کد مقاله | کد نشریه | سال انتشار | مقاله انگلیسی | نسخه تمام متن |
|---|---|---|---|---|
| 1393472 | 1501213 | 2008 | 12 صفحه PDF | دانلود رایگان |
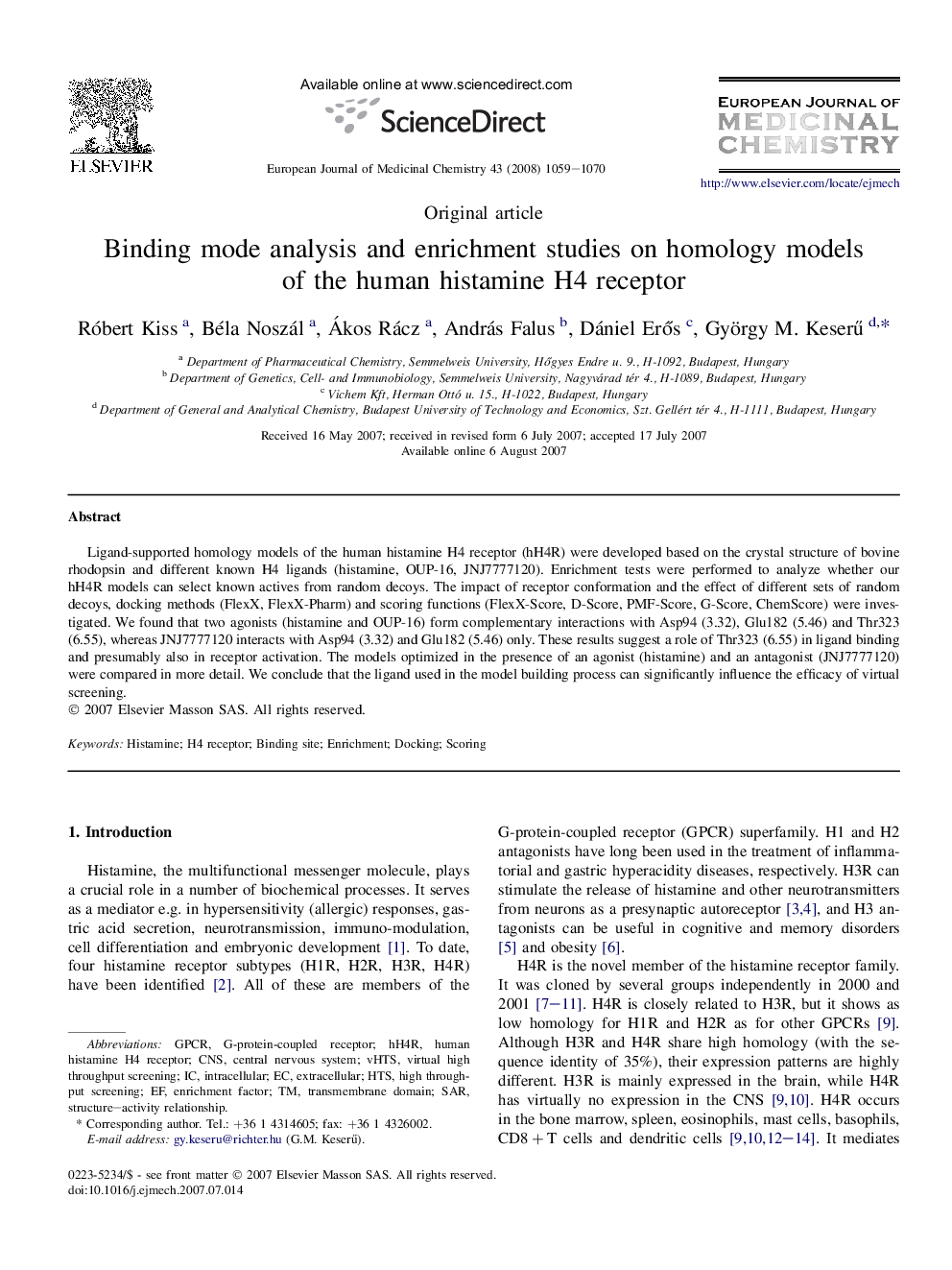
Ligand-supported homology models of the human histamine H4 receptor (hH4R) were developed based on the crystal structure of bovine rhodopsin and different known H4 ligands (histamine, OUP-16, JNJ7777120). Enrichment tests were performed to analyze whether our hH4R models can select known actives from random decoys. The impact of receptor conformation and the effect of different sets of random decoys, docking methods (FlexX, FlexX-Pharm) and scoring functions (FlexX-Score, D-Score, PMF-Score, G-Score, ChemScore) were investigated. We found that two agonists (histamine and OUP-16) form complementary interactions with Asp94 (3.32), Glu182 (5.46) and Thr323 (6.55), whereas JNJ7777120 interacts with Asp94 (3.32) and Glu182 (5.46) only. These results suggest a role of Thr323 (6.55) in ligand binding and presumably also in receptor activation. The models optimized in the presence of an agonist (histamine) and an antagonist (JNJ7777120) were compared in more detail. We conclude that the ligand used in the model building process can significantly influence the efficacy of virtual screening.
Ligand-supported homology models of the human histamine H4 receptor were developed. Our results suggest a different binding mode of histamine than proposed previously. Enrichment studies demonstrated the suitability of the models for virtual screening. Figure optionsDownload as PowerPoint slide
Journal: European Journal of Medicinal Chemistry - Volume 43, Issue 5, May 2008, Pages 1059–1070