| کد مقاله | کد نشریه | سال انتشار | مقاله انگلیسی | نسخه تمام متن |
|---|---|---|---|---|
| 8320289 | 1539357 | 2010 | 10 صفحه PDF | دانلود رایگان |
عنوان انگلیسی مقاله ISI
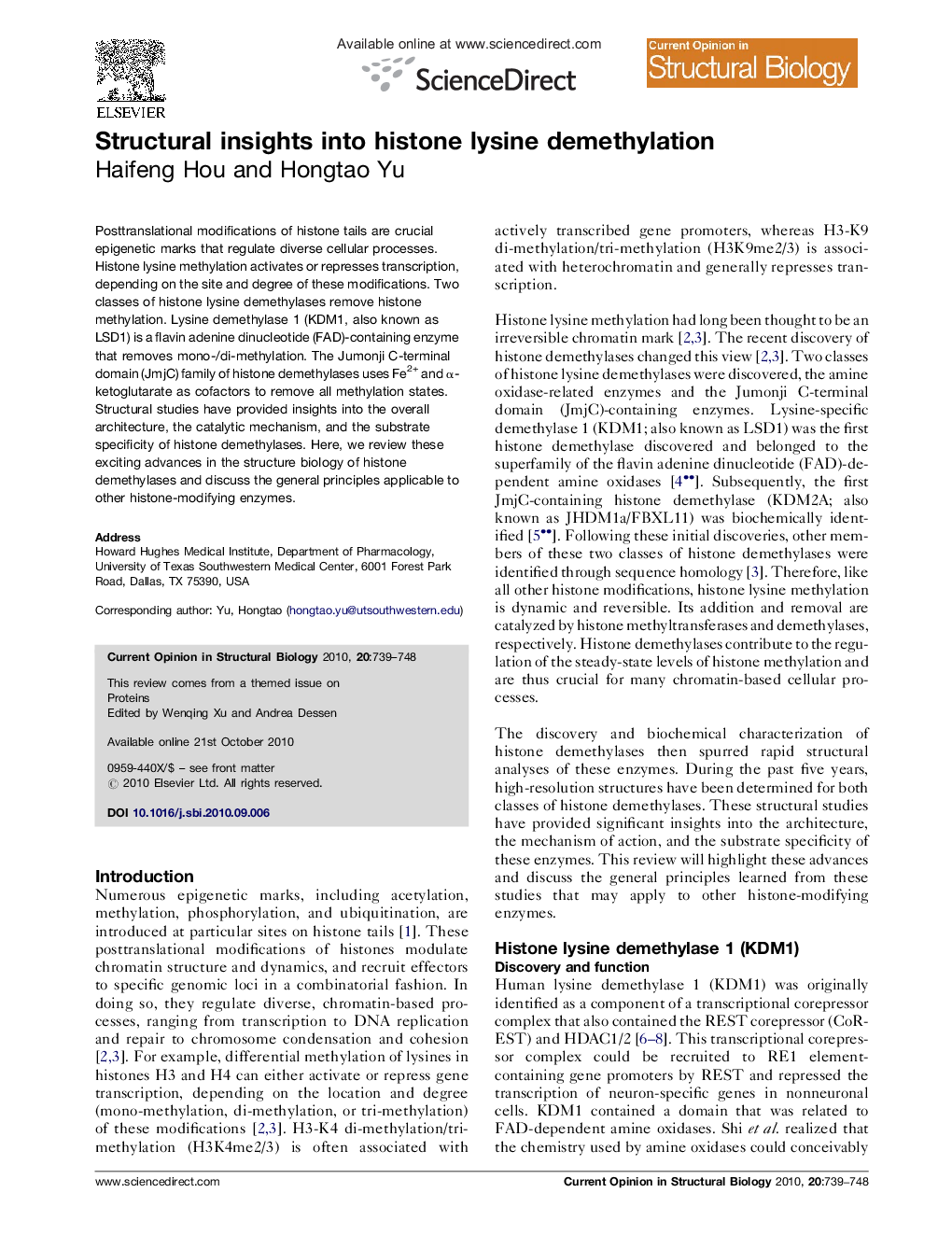
Structural insights into histone lysine demethylation
دانلود مقاله + سفارش ترجمه
دانلود مقاله ISI انگلیسی
رایگان برای ایرانیان
موضوعات مرتبط
علوم زیستی و بیوفناوری
بیوشیمی، ژنتیک و زیست شناسی مولکولی
زیست شیمی
پیش نمایش صفحه اول مقاله
چکیده انگلیسی
Posttranslational modifications of histone tails are crucial epigenetic marks that regulate diverse cellular processes. Histone lysine methylation activates or represses transcription, depending on the site and degree of these modifications. Two classes of histone lysine demethylases remove histone methylation. Lysine demethylase 1 (KDM1, also known as LSD1) is a flavin adenine dinucleotide (FAD)-containing enzyme that removes mono-/di-methylation. The Jumonji C-terminal domain (JmjC) family of histone demethylases uses Fe2+ and α-ketoglutarate as cofactors to remove all methylation states. Structural studies have provided insights into the overall architecture, the catalytic mechanism, and the substrate specificity of histone demethylases. Here, we review these exciting advances in the structure biology of histone demethylases and discuss the general principles applicable to other histone-modifying enzymes.
ناشر
Database: Elsevier - ScienceDirect (ساینس دایرکت)
Journal: Current Opinion in Structural Biology - Volume 20, Issue 6, December 2010, Pages 739-748
Journal: Current Opinion in Structural Biology - Volume 20, Issue 6, December 2010, Pages 739-748
نویسندگان
Haifeng Hou, Hongtao Yu,