| کد مقاله | کد نشریه | سال انتشار | مقاله انگلیسی | نسخه تمام متن |
|---|---|---|---|---|
| 5131053 | 1490862 | 2017 | 7 صفحه PDF | دانلود رایگان |
- A real-time and one-step fluorescent method for detecting methyltransferase activity is proposed.
- The approach is based on enzyme-free entropy-driven toehold-mediated hairpin displacement.
- High sensitivity with a low detection limit of 0.01 UÂ mLâ1 is achieved.
- The application of high-throughput screening drugs as methyltransferase inhibitors is demonstrated.
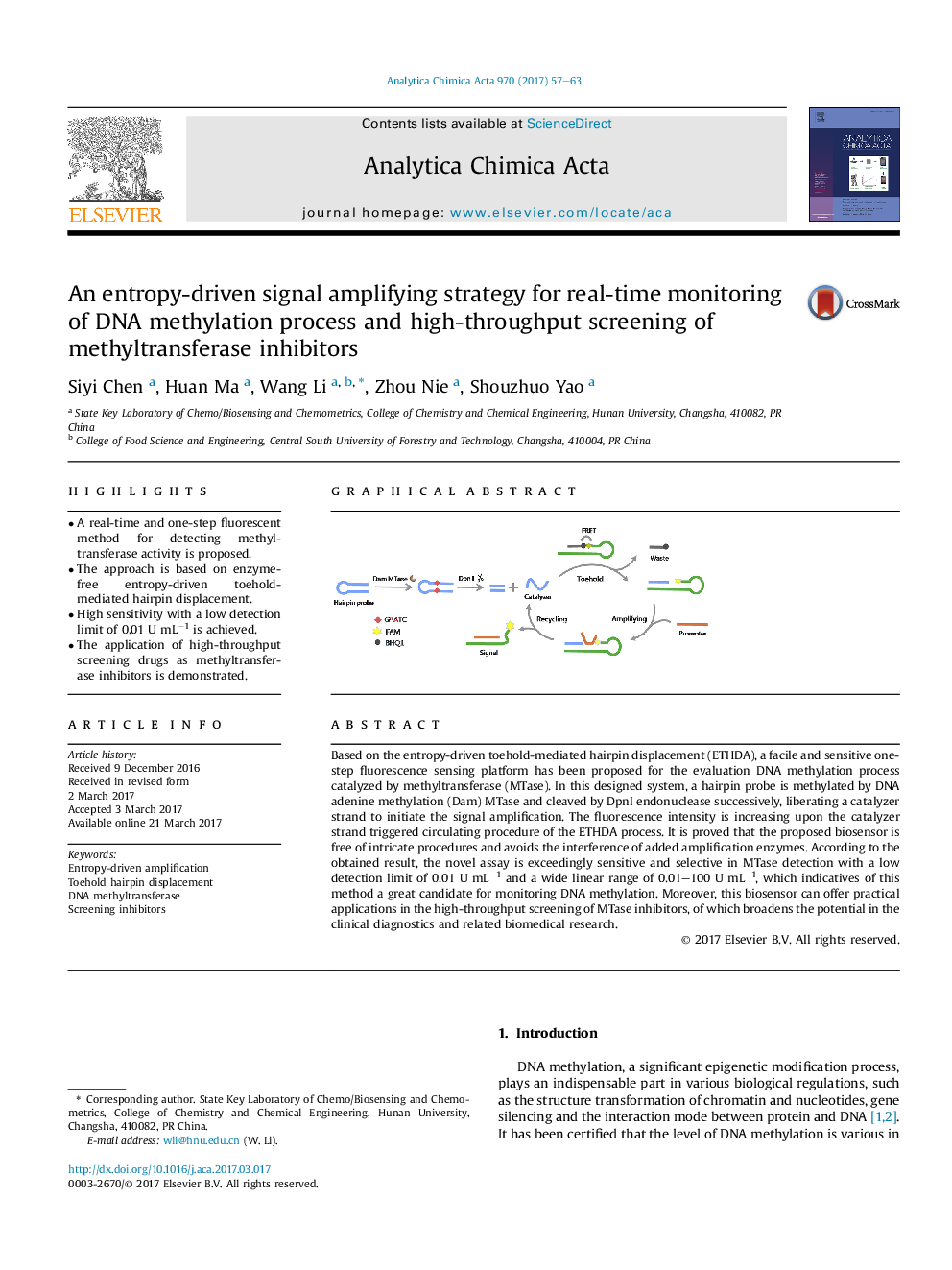
Based on the entropy-driven toehold-mediated hairpin displacement (ETHDA), a facile and sensitive one-step fluorescence sensing platform has been proposed for the evaluation DNA methylation process catalyzed by methyltransferase (MTase). In this designed system, a hairpin probe is methylated by DNA adenine methylation (Dam) MTase and cleaved by DpnI endonuclease successively, liberating a catalyzer strand to initiate the signal amplification. The fluorescence intensity is increasing upon the catalyzer strand triggered circulating procedure of the ETHDA process. It is proved that the proposed biosensor is free of intricate procedures and avoids the interference of added amplification enzymes. According to the obtained result, the novel assay is exceedingly sensitive and selective in MTase detection with a low detection limit of 0.01 UÂ mLâ1 and a wide linear range of 0.01-100 UÂ mLâ1, which indicatives of this method a great candidate for monitoring DNA methylation. Moreover, this biosensor can offer practical applications in the high-throughput screening of MTase inhibitors, of which broadens the potential in the clinical diagnostics and related biomedical research.
This one-step fluorescence sensing platform takes advantage of the entropy-driven toehold-mediated hairpin displacement amplification (ETHDA) for real-time monitoring of DNA methylation process and high-throughput screening of methyltransferase inhibitors.149
Journal: Analytica Chimica Acta - Volume 970, 1 June 2017, Pages 57-63