| کد مقاله | کد نشریه | سال انتشار | مقاله انگلیسی | نسخه تمام متن |
|---|---|---|---|---|
| 1162741 | 1490898 | 2016 | 7 صفحه PDF | دانلود رایگان |
• An alternative method for species identification of wild mushroom was proposed.
• Rapid and reliable identification was achieved by use of MALDI-TOF MS.
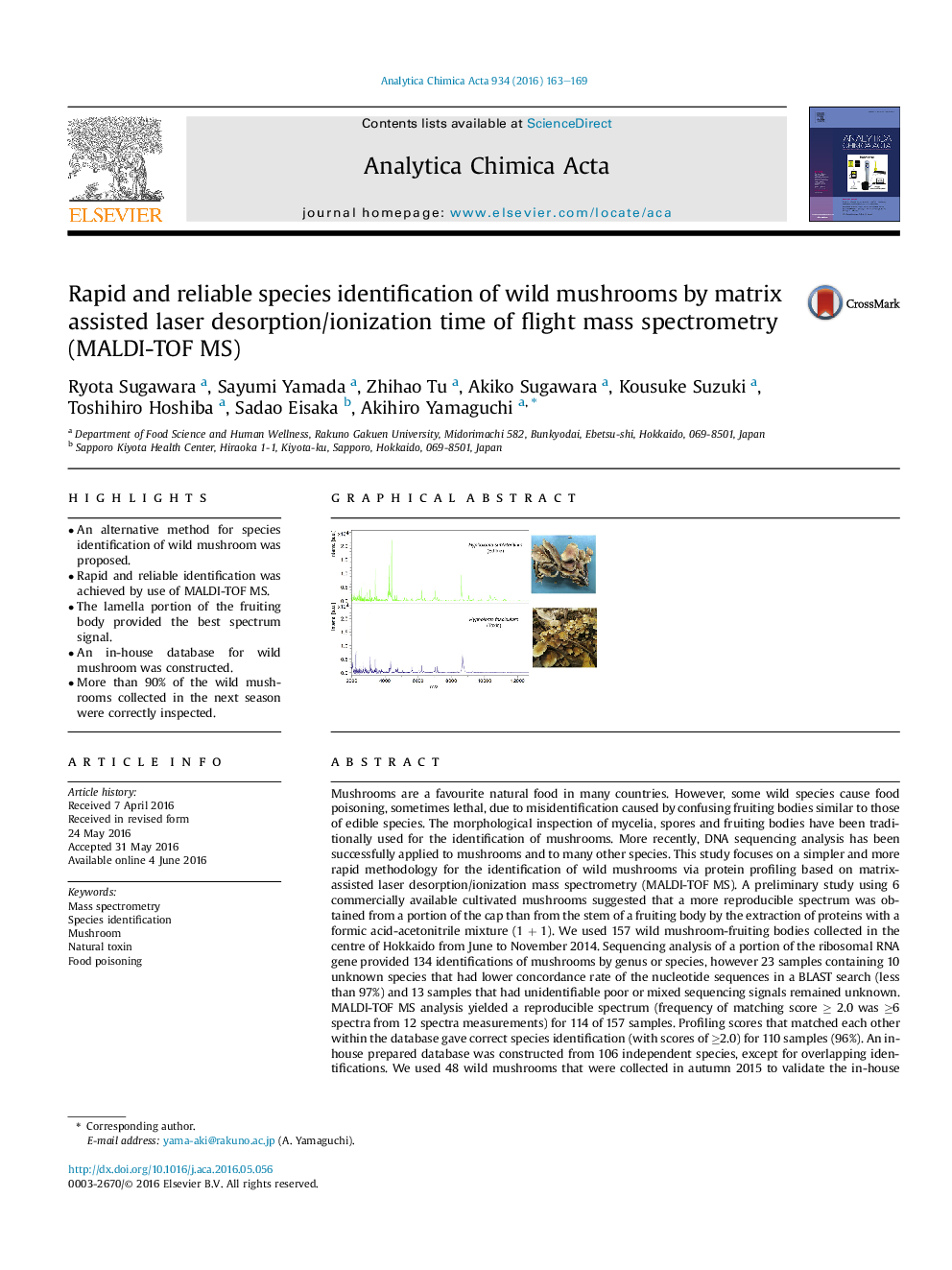
• The lamella portion of the fruiting body provided the best spectrum signal.
• An in-house database for wild mushroom was constructed.
• More than 90% of the wild mushrooms collected in the next season were correctly inspected.
Mushrooms are a favourite natural food in many countries. However, some wild species cause food poisoning, sometimes lethal, due to misidentification caused by confusing fruiting bodies similar to those of edible species. The morphological inspection of mycelia, spores and fruiting bodies have been traditionally used for the identification of mushrooms. More recently, DNA sequencing analysis has been successfully applied to mushrooms and to many other species. This study focuses on a simpler and more rapid methodology for the identification of wild mushrooms via protein profiling based on matrix-assisted laser desorption/ionization mass spectrometry (MALDI-TOF MS). A preliminary study using 6 commercially available cultivated mushrooms suggested that a more reproducible spectrum was obtained from a portion of the cap than from the stem of a fruiting body by the extraction of proteins with a formic acid-acetonitrile mixture (1 + 1). We used 157 wild mushroom-fruiting bodies collected in the centre of Hokkaido from June to November 2014. Sequencing analysis of a portion of the ribosomal RNA gene provided 134 identifications of mushrooms by genus or species, however 23 samples containing 10 unknown species that had lower concordance rate of the nucleotide sequences in a BLAST search (less than 97%) and 13 samples that had unidentifiable poor or mixed sequencing signals remained unknown. MALDI-TOF MS analysis yielded a reproducible spectrum (frequency of matching score ≥ 2.0 was ≥6 spectra from 12 spectra measurements) for 114 of 157 samples. Profiling scores that matched each other within the database gave correct species identification (with scores of ≥2.0) for 110 samples (96%). An in-house prepared database was constructed from 106 independent species, except for overlapping identifications. We used 48 wild mushrooms that were collected in autumn 2015 to validate the in-house database. As a result, 21 mushrooms were identified at the species level with scores ≥2.0 and 5 mushrooms at the genus level with scores ≥1.7, although the signals of 2 mushrooms were insufficient for analysis. The remaining 20 samples were recognized as “unreliable identification” with scores <1.7. Subsequent DNA analysis confirmed that the correct species or genus identifications were achieved by MALDI-TOF MS for the 26 former samples, whereas the 18 mushrooms with poorly matched scores were species that were not included in the database. Thus, the proposed MALDI-TOF MS coupled with our database could be a powerful tool for the rapid and reliable identification of mushrooms; however, continuous updating of the database is necessary to enrich it with more abundant species.
Figure optionsDownload as PowerPoint slide
Journal: Analytica Chimica Acta - Volume 934, 31 August 2016, Pages 163–169